
Revision Notes on Biotechnology Principles and Processes NEET 2025 - Free PDF Download
Biotechnology Process and Principles is one of the most important chapters for NEET preparation. It is a very crucial chapter since it introduces the students to different concepts of biotechnology, such as genetic engineering, bioprocess engineering, Recombinant DNA Technology and much more.
Students can develop a detailed foundation about the topic. Biotechnology is a scientific field of study that revolves around the development of different technologies that are supposedly targeted towards the development of human beings. In case there are any doubts regarding the important topics of the chapter, refer to the Biotechnology Principles and Processes Class 12 notes. Students can take help from these notes that have been prepared by subject matter experts at Vedantu.
Access NEET Revision Notes Biology Biotechnology: Principles and Processes
Introduction
Biotechnology can be defined as the use of microorganisms, animals, or plant cells, or their products, to produce a variety of products and services that are valuable to humans on an industrial scale.
Traditional biotechnology relied on microorganisms' inherent abilities.
For example, Citric acid is produced by Penicillium notatum, while Penicillin is produced by Penicillium notatum.
Recombinant DNA technology is the heart of new biotechnology. A human gene that produces insulin, for example, has been transferred and expressed in bacteria such as Escherichia coli.
Microbiology, biochemistry, tissue culture, chemical engineering and genetic engineering, molecular biology, and immunology are all used in modern biotechnology to create a variety of valuable products.
Genetic Engineering:
One type of biotechnology that involves DNA manipulation is genetic engineering (also known as recombinant DNA technology or gene splicing). It is concerned with the isolation of relevant genes from various sources as well as the production of novel DNA combinations (recombinant DNA) for genotype repair, improvement, perfection, and matching.
Genetic engineering is a technique for artificially and purposefully altering DNA (genes) to meet human requirements.
Breaking a DNA molecule at two specified locations with the help of restriction endonuclease to extract a specific DNA segment and then inserting it in another DNA molecule at a desired position is used in genetic engineering to manipulate it.
Recombinant DNA is the new DNA molecule, and genetic engineering is the technique. The goal of genetic engineering is to add, remove, or repair a piece of genetic material.
With the help of the - phage, Paul Bergh (Father of Genetic Engineering) introduced the gene of the SV 40 virus (Simian virus) into E.coli. (Nobel Prize in Physics, 1980)
The concept of genetic engineering was born out of two major breakthroughs in bacterial research. These had been –
Extrachromosomal DNA segments termed plasmids are found in bacterial cells and reproduce alongside the bacterium's chromosomal DNA.
Restriction endonucleases are enzymes that cleave DNA at a specified site. As a result, these enzymes are known as molecular scissors.
Microbiology, biochemistry, tissue culture, and chemical engineering are used to create a variety of valuable products.
Tools and Techniques of Genetic Engineering
Restriction Enzymes
In genetic engineering, a number of different enzymes are used.
Lysing enzymes are employed to open cells in order to obtain DNA for genetic experiments. Lysozyme is often used to break down bacterial cell walls.
Exonucleases, endonucleases, and restriction endonucleases are the three types of cleaving enzymes.
Exonucleases cut off nucleotides from 5' or 3' ends of DNA molecules.
Endonucleases break DNA duplexes at any point except the end.
Restriction Endonucleases cleave DNA duplexes at specific points in such a way that they come to possess short single-stranded free ends.
Arber, Smith, and Nathans identified the EcoRI restriction enzyme (1978 Nobel prize). These enzymes are found in many microorganisms, and in addition to cleavage, some restriction endonucleases can also modify DNA.
Restriction enzymes are utilised in recombinant DNA technology because they can recognise and cleave certain DNA sequences in vitro, which are typically 4 to 8 nucleotides long. The restriction site is a 4 to 8 nucleotide region that is frequently palindromic, meaning that the DNA sequence is the same on both DNA strands when read in the 5'-3' orientation.
As a result, the DNA fragments produced by these enzymes contain a brief single-stranded overhang at each end, which is referred to as sticky or cohesive ends because base pairing between them can reassemble the DNA molecule.
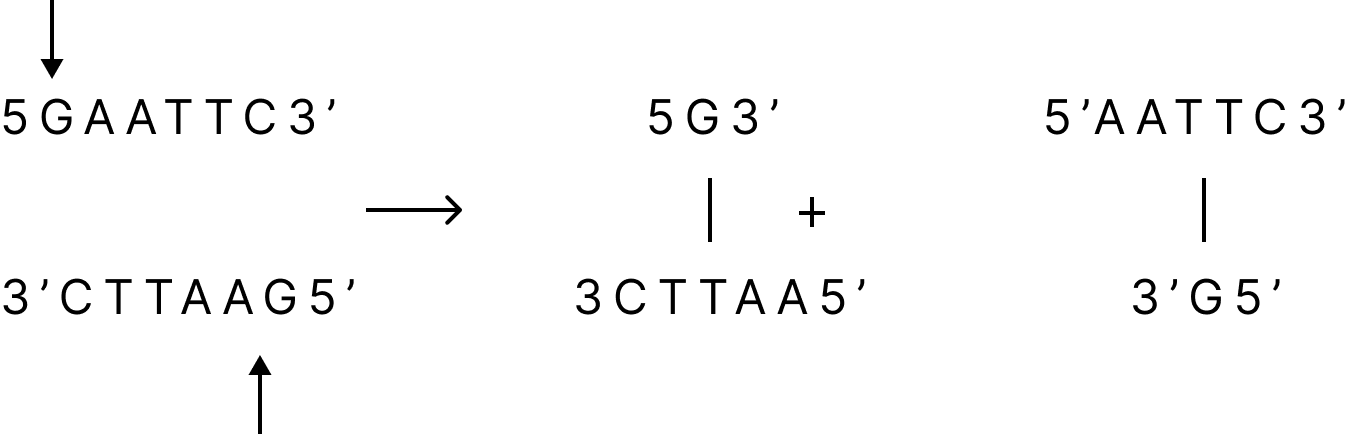
Restriction Enzymes generating Sticky Ends
A recombinant DNA molecule can be created by cutting two distinct DNA samples with the same restriction enzyme and combining the fragments together.
Some enzymes, on the other hand, break both strands of DNA at the same nucleotide position, usually in the middle of the recognition sequence, resulting in a blunt or flush termination.
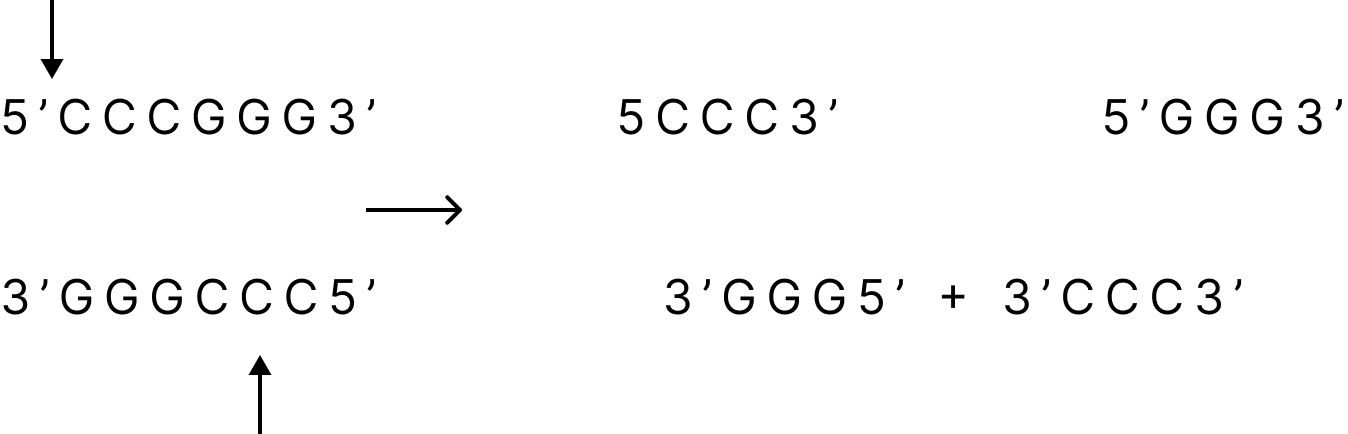
Restriction Enzymes Generating Blunt Ends
These fragments can be separated using a process called gel electrophoresis, which takes use of the fact that these molecules have charged groups that lead them to move through a matrix under an electric field.
Agarose, a natural polymer produced from seaweeds, is the most widely utilised medium in laboratories for bacteria culture, gel electrophoresis, etc.
Only after dyeing the DNA with a chemical called ethidium bromide and then exposing it to UV rays can the separated DNA fragments be seen.
The divided bands of DNA are cut out of the agarose gel portion and extracted. This process is referred to as Elution.
Enzymes that synthesise: These enzymes are used to make new DNA strands that are complementary to a DNA or RNA template.
Reverse Transcriptases and DNA Polymerases are the Two Types:
Reverse Transcriptases help in the synthesis of complementary DNA strands on RNA templates;
DNA Polymerases help in the synthesis of complementary DNA strands on DNA templates.
Joining Enzymes: These enzymes aid in the recombination of DNA fragments. DNA ligase from Escherichia coli, for example, is used to connect DNA fragments. Molecular glues are enzymes that join molecules together.
Alkaline Phosphatases: These enzymes prevent recircularization by removing a phosphate group from the 5' end of linearized circular DNA.
Cloning Vector
Vehicle or vector DNA is the DNA that is employed as a carrier for introducing a segment of foreign DNA into a suitable host. Plasmids and bacteriophages can reproduce in bacterial cells without the help of chromosomal DNA.
The following are the characteristics that must be present in order for cloning into a vector to be possible.
Replication's starting point (Ori): This is the sequence in which replication begins. For the vector DNA to divide within the host cells to produce its more copies, it needs an origin of replication (ori). This sequence is also in charge of regulating the quantity of copies of linked DNA. This means that any foreign DNA introduced into the vector will be replicated during the replication phase.
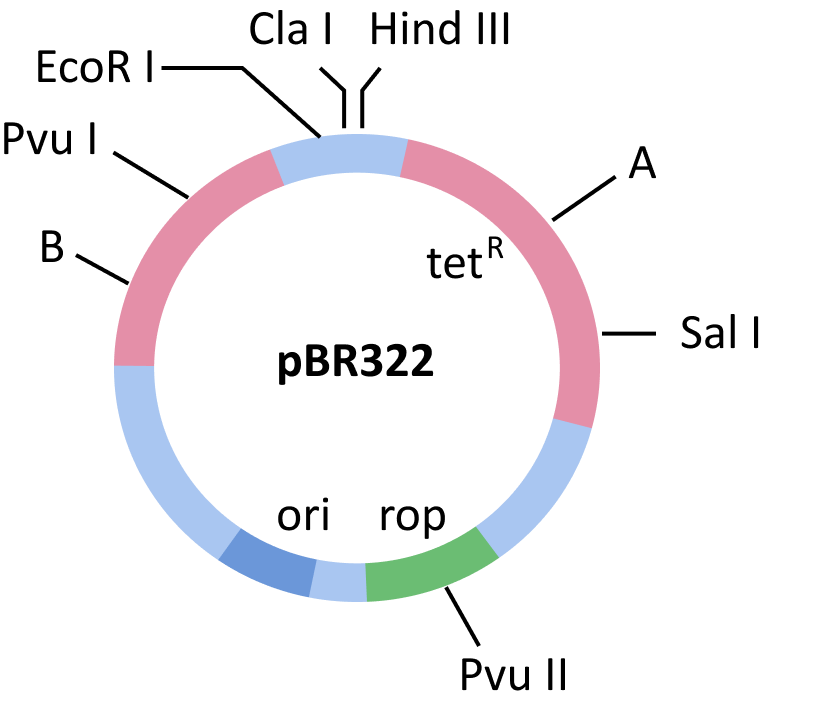
Cloning Vector pBR322.
Selectable Marker: The vector also requires a selectable marker in addition to 'ori.' Antibiotic resistance genes, such as ampicillin, chloramphenicol, tetracycline, or kanamycin resistance genes, are usually considered valuable selection markers for E. coli.
Cloning Sites: The vector requires recognition sites for commonly used restriction enzymes in order to link the alien DNA. A restriction site located in one of the two antibiotic resistance genes is used to ligate foreign DNA. For example, with the vector pBR322, you can ligate a foreign DNA at the BamHI site of the tetracycline resistance gene. Due to the inclusion of foreign DNA, the recombinant plasmids will lose their tetracycline resistance (insertional inactivation). It may now be distinguished from non-recombinant transformants by plating transformants on ampicillin-containing media. After growing on ampicillin-containing media, the transformants (plasmid transfer) are moved to a tetracycline-containing medium. In the ampicillin-containing medium, the recombinants will grow, but not in tetracycline-containing medium. Non-recombinants, on the other hand, will grow on a medium containing both antibiotics. One antibiotic resistance gene aids in the selection of transformants in this circumstance.
Selection of recombinants is a difficult method due to antibiotic inactivation, as it necessitates simultaneous plating on two plates with different antibiotics. As a result, alternative selectable markers that distinguish recombinants from non-recombinants based on their ability to produce colour in the presence of a chromogenic substrate have been devised. Insertional inactivation is a technique that involves inserting recombinant DNA into an enzyme's coding sequence. If the plasmid in the bacterium does not include an insert, the presence of the chromogenic substrate X-gal (5-bromo-4,chloro—D galactopyranoside) results in blue colonies. The presence of an insert causes the -galactosidase (reporter enzyme) to be inactivated, and the colonies do not produce any colour, identifying them as recombinant colonies.
Vectors for Cloning Genes in Plants and Animals : Agrobacterium tumefaciens uses a fragment of DNA called T-DNA to turn normal plant cells into tumours. In animals, retroviruses have the power to change healthy cells into malignant ones. Pathogens' tools for delivering genes to their eukaryotic hosts have been transformed into useful vectors for delivering genes of interest to humans, thanks to a greater understanding of the art of delivering genes by pathogens in their eukaryotic hosts.
Plasmids: Extra chromosomal DNA segments found in bacteria that may replicate independently are known as plasmids. Restriction enzymes and DNA ligase can be used to extract plasmids from bacteria and join them with appropriate DNA segments. Recombinant plasmids, hybrid plasmids, and chimeric plasmids are plasmids that have the DNA of another organism integrated into them.
pBR vector plasmids, for example (named after the discoverer Bolivar and Rodriguez, pUC vector plasmid University of California)
Viruses: Certain viruses have DNA that can be used as a vehicle DNA. The genes for galactosidase were transferred from Escherichia coli to human cells using a bacteriophage (bacterial virus). Lambda phage (phage) was utilised to transfer lac genes from E. coli to tomato haploid callus.
Note:
Vector Type | Insert Size (in kb) |
Plasmid | 0.5-8 |
Bacteriophage lambda | 9-23 |
Cosmid | 30-45 |
BAC | 50-300 |
YAC | 1000-2500 |
Passenger DNA: It is the DNA that is combined with the vehicle DNA and transported from one organism to another. Complementary, synthetic, or random DNA can be used as a passenger.
Complementary DNA (cDNA) is created using reverse transcriptase and the required nucleotides on an mRNA template.
Synthetic DNA (sDNA) is created on a DNA template with the help of DNA polymerase.
Random DNA refers to tiny pieces created when restriction endonucleases are used to break a chromosome.
Process of Recombinant DNA Technology
A recombinant DNA molecule is created when two or more DNA segments from different organisms are joined together. A recombinant DNA molecule is a carrier (such as a plasmid, phage, or virus) into which the desired DNA fragment has been inserted to allow it to be cloned in a suitable host.
Recombinant Dna Molecules are Created With One of Three Goals in Mind:
To get a lot of copies of a certain DNA segment,
To recover a large amount of the protein generated by the gene in question.
To integrate the gene in issue into a target organism's genome, where it can express itself.
Genetic engineering was used to develop this approach. First, the desired gene is isolated from any creature and then transferred and expressed into the organism of choice. These are known as transgenic organisms. In order to obtain novel therapeutic proteins, transgenic microbes are created.
Human insulin, for example, is commercially produced from a transgenic E.coli strain.
Transgenic animal cell lines and transgenic plants are also used to make a variety of useful recombinant proteins.
At the same time, non-transgenic cells or organisms could not manufacture a number of these important medical proteins on a commercial scale. Recombinant proteins are proteins created by transgenes. Recombinant genes of this type are used to create a variety of products.
Application of Recombinant DNA Technology
The recombinant DNA technology can be used in the following ways:
It can be used to decipher molecular events in biological processes like cell differentiation and ageing. The same method can be used to create precise gene maps.
Useful chemical compounds can be created inexpensively and efficiently in the biochemical and pharmaceutical industries by engineering genes.
Transgenic plant production.
Microorganisms that have been genetically manipulated.
The recombinant DNA technique entails a series of steps that must be completed in a certain order, such as
Isolation of a particular type of genetic material
DNA is cut at certain spots.
The polymerase chain reaction is used to amplify a gene of interest.
Recombinant DNA is inserted into the host cell/organism.
Getting a hold of the foreign gene product.
Processes that occur downstream.
Isolation of a Specific Genetic Material (DNA)
Isolation is the process of removing plasmids or genomic DNA from cells.
Isolation normally entails rupturing the cell membrane (and perhaps the nuclear membrane) as well as the cell wall (plant cells). DNA is generated by using enzymes such as lysozyme (bacteria), cellulase (plant cells), and chitinase (animal tissue) to treat bacterial cells, plant cells, or animal tissue (fungus). Ribonuclease can be used to remove RNA, while protease can be used to remove proteins. Other compounds are eliminated with the use of appropriate treatments. Following the addition of cold ethanol, the pure DNA precipitates out.
Cutting of DNA at Specific Locations
Only after DNA from an organism has been broken into smaller molecules can it be prepared for cloning.
DNA cleavage is made feasible by the restriction of endonucleases.
To monitor the progress of restriction enzyme digestion, agarose gel electrophoresis is used. The procedure is carried out once again with the vector DNA. The cut out gene of interest from the source DNA and the cut vector with space are mixed and ligase is added after the source and vector DNA have been cut using a specific restriction enzyme. DNA ligase is a protein that binds two pieces of DNA together. Recombinant DNA is created as a result of this process.
Amplification of Gene of Interest using Polymerase Chain Reaction (PCR)
It is a laboratory method used to make many copies of a specific piece of DNA from a sample that contains very tiny amounts of that DNA. Using two sets of primers and the enzyme DNA polymerase, multiple copies of the gene (or DNA) of interest are generated in vitro.
The enzyme uses the nucleotides in the process and the genomic DNA as a template to lengthen the primers.
The PCR Cycle has Following Three Steps:
1. Denaturation: In this phase, double-stranded DNA is heated to 94°C for two minutes to separate the two strands. The hydrogen bonds between the base pairs break, causing the two strands to split.
2. Annealing:The process of annealing involves the binding of primers, which are 20 base pairs (bp) long, to the 3' end of single strands of DNA. In order for the primers to anneal to the complementary sequences of the isolated single strands of DNA, the reaction temperature is decreased to 56°C.
3. Extension: In this step, the temperature of the reaction is raised to 72°C. The primers are extended by the enzyme Taq Polymerase. The enzyme adds nucleotides to the 3' end of the primers and extends them until a DNA strand which is complementary to the template DNA is formed.
Thus, one PCR cycle is completed and the amount of DNA doubles and so on.
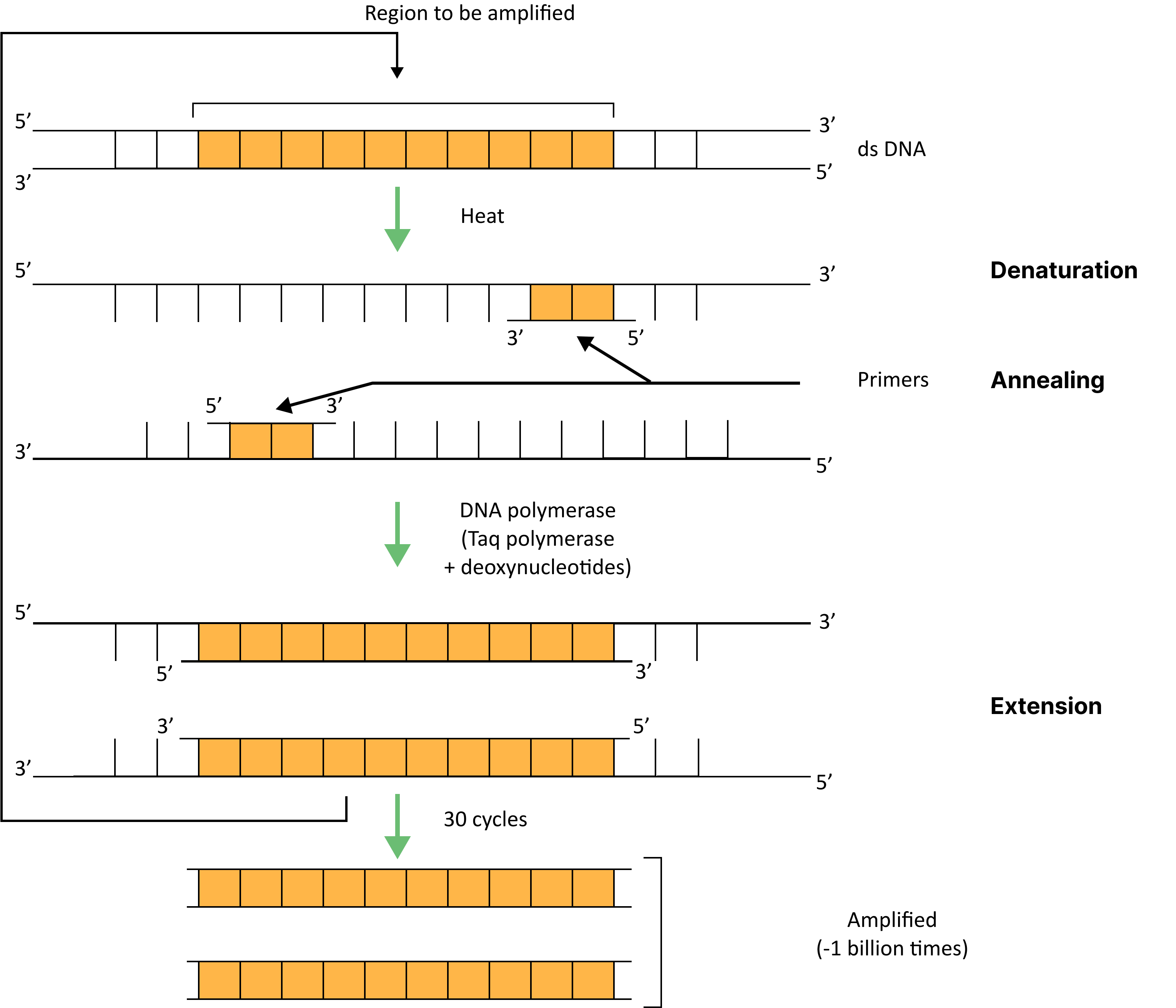
Polymerase Chain Reaction
Insertion of Recombinant DNA Into the Host Cell/Organism
Once connected to a DNA vector, the desired DNA sequence must be transmitted to an appropriate host. The introduction of foreign DNA into a recipient cell is known as transformation. Transfection refers to the process of infecting a cell with DNA from a virus. There are a few different ways to get the ligated DNA into the recipient cells. The host is commonly Escherichia coli, and E. coli transformation is a necessary step in this research. DNA can be transferred into cells of higher eukaryotes via a variety of methods.
Obtaining the Foreign Gene Product
Recombinant DNA technology's ultimate goal is to create a desirable protein. As a result, the expression of recombinant DNA is required. Under the right circumstances, the foreign gene is expressed. The cultures can be utilised to extract the necessary protein, which can subsequently be purified using various separation processes.
It was necessary to construct bioreactors that could process enormous volumes of culture (100-1000 litres) in order to manufacture in large quantities. Bioreactors are tanks in which raw materials are biologically transformed into specialised products, individual enzymes, and so on, employing microbial plant, animal, or human cells. A bioreactor provides the best circumstances for producing the desired product by allowing for optimal growth (temperature, pH, substrate, salts, vitamins, oxygen).
To aid in the mixing of the reactor contents, stirred-tank reactors are usually cylindrical or have a curved base. The stirrer ensures that the bioreactor is evenly mixed and that oxygen is available at all times. Air can also be bubbled through the reactor as an alternative. The bioreactor is equipped with an agitator system, an oxygen delivery system, a foam control system, a temperature control system, a pH control system, and sampling parts that allow tiny amounts of culture to be removed at regular intervals.
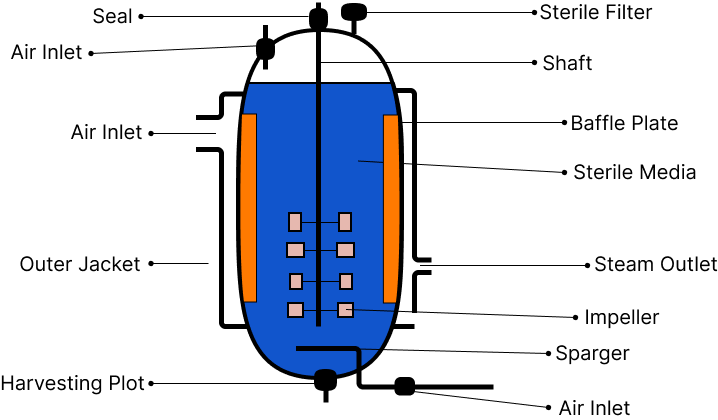
Simple Stirred-Tank Bioreactor
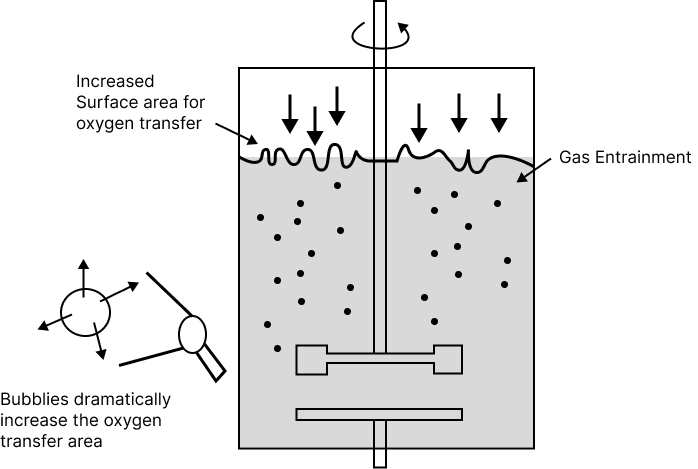
Sparged Stirred Tank Bioreactor
Microorganisms Can Be Grown in Bioreactors in Two Ways:
Support Growth System: In this method, microorganisms are grown as a thin layer or film in a solid medium.
Suspended Growth System: By suspending cells or mycelia in the liquid medium is called a suspended growth system.
Manufacturing Unit: During the designing of a bioreactor for the process, often a very large-sized unit is used so that it accommodates a huge amount of medium.
Downstream Process:
After the biosynthetic stage is completed, the product must go through a series of processes before it can be sold as a finished product. Separation and purification are among the procedures, which are referred to as downstream processing. Preservatives must be included in the product's formulation.
Key Points to Remember
Biotechnology deals with the large-scale production and marketing of products and processes using live organisms, cells or enzymes.
The PCR cycle has the Following Three Steps:
In the Denaturation phase, double-stranded DNA is heated to 94°C for two minutes to separate the two strands.
In Annealing, the process of annealing involves the binding of primers, which are 20 base pairs (bp) long, to the 3' end of single strands of DNA.
Extension, in this step, the temperature of the reaction is raised to 72°C. The primers are extended by the enzyme Taq Polymerase. Thus, one PCR cycle is completed and the amount of DNA doubles and so on.
Modern biotechnology uses genetically modified organisms to alter the chemistry of DNA and construct recombinant DNA. This process is known as recombinant DNA technology or genetic engineering.
It involves the use of restriction endonucleases, DNA ligase, appropriate plasmid or viral vectors to isolate the foreign DNA into host organisms, expression of the foreign gene, and purification of the gene product.
Large-scale production involves the use of bioreactors.
Importance of NEET Biology Biotechnology Principles and Processes Notes
Modern Biotechnology has a very important chapter for students of NEET. With this chapter, students will be introduced to the concept of Genetic Engineering or the manipulation of an organism's gene. They can also learn how a sterile environment is maintained through chemical engineering for the growth of microbes. The concepts are well explained by the subject matter experts at Vedantu.
Students will also be introduced to other important concepts and processes of Biotechnology such as the Development of rDNA or Recombinant DNA and the cloning of any desired gene. They can learn about the process of transferring cloned genes to any organism.
The introduction to different tools used for Genetic Engineering will help students understand the process. They will get to know about recognition sequences, endonucleases, Isolation and Separation of DNA fragments with electrophoresis, elution, visualisation, and much more. Biotechnology Principles and Processes notes are definitely helpful for students who want to get a detailed view of the chapter and what it entails.
Benefits of Biotechnology Principles and Processes Notes PDF
After downloading the PDF files of NEET revision notes for the Biotechnology Principles and Processes chapter, students can gain a plethora of benefits in the following ways:
The experts at Vedantu have carefully detailed all the processes and different principles related to biotechnology such as genetic manipulation, recombinant DNA technology, and other important processes. Students can get a detailed account of these topics and build their conceptual foundation using the knowledge.
Students will get to learn what is the origin of replication and the scope of biotechnology in different fields including medicine and health sectors.
Students can understand what cloning vectors are and what the important parts of the process of recombinant DNA technology are. They will also learn what a bioreactor is, what are its types and the different parts that it has.
Students can prepare well for their NEET entrance examination with the help of Biotechnology Principles and Processes NEET notes. Doubt clarification and answering questions from the chapter becomes really easy with these notes.
Download Biotechnology Principles and Processes Notes PDF
Want to score high marks in your NEET entrance exams to secure a place in the top medical colleges and institutions? Well, download the Biotechnology Principles and Processes notes PDF from Vedantu. Downloading the notes will ensure that you are able to prepare for your exams in the best way. Every student visiting the official website of Vedantu can access the PDFs and download the notes.
Other Important Links
Other Important Links for NEET Biotechnology Principles and Processes |
NEET Biotechnology Principles and Processes Important Chapter |
NEET Biotechnology Principles and Processes Important Questions |
NEET Biology Revision Notes - Chapter Pages
NEET Biology Chapter-wise Revision Notes | |
Biotechnology Principles and Processes Notes | |
FAQs on Revision Notes on Biotechnology Principles and Processes NEET 2025
1. What is bioprocess engineering?
It is a process that involves the design of necessary proteins that are extracted from a huge amount of different proteins. The products that are extracted will go through different subjective procedures and quality checks before being launched for use. Bioprocess engineering is mostly used for the production of antibiotics, drugs, and vaccines.
2. What are the different tools of recombinant DNA technology?
The different tools of recombinant DNA technology include: Restriction enzymes, Polymerase enzymes, Vectors, and Competent host organism.
3. What is a cloning vector?
Cloning vectors can be considered as DNA molecules that have the primary function of carrying different foreign fragments of DNA into any host cell.
4. Define gel electrophoresis?
It is the process of migrating the negatively charged DNA towards any electrode that is positively charged using a porous polymer gel matrix and passing electric current in the electric field. The fragments of DNA start moving around in the gel and hence will resolve. Agarose is a very common gel matrix often used for the process of DNA electrophoresis.


















